How to Select a Target Protein for Computational Drug Discovery
Introduction

Selecting the right target protein is a critical first step in computational drug discovery. A well-chosen target increases the likelihood that virtual screening, molecular docking, and molecular dynamics simulations will yield meaningful and biologically relevant results.
This tutorial introduces practical and beginner-friendly approaches to identifying and validating target proteins. The focus is on conceptual understanding, not software-specific instructions. Future tutorials from Idgence Research will explore each method in detail with hands-on examples.
1. Start with a Disease-Oriented Literature Review
A common and effective strategy is to begin with a target disease and identify proteins known to play important roles in its pathology.
Example: Alzheimer’s Disease
- Search PubMed or Google Scholar
- Use clear and focused keywords such as:
- Molecular targets of Alzheimer’s disease
- Novel therapeutic targets for Alzheimer’s disease
- Kinases involved in neurodegeneration
- Prioritize review articles authored by recognized experts. Reviews summarize:
- Established targets
- Emerging targets
- Known druggable pathways
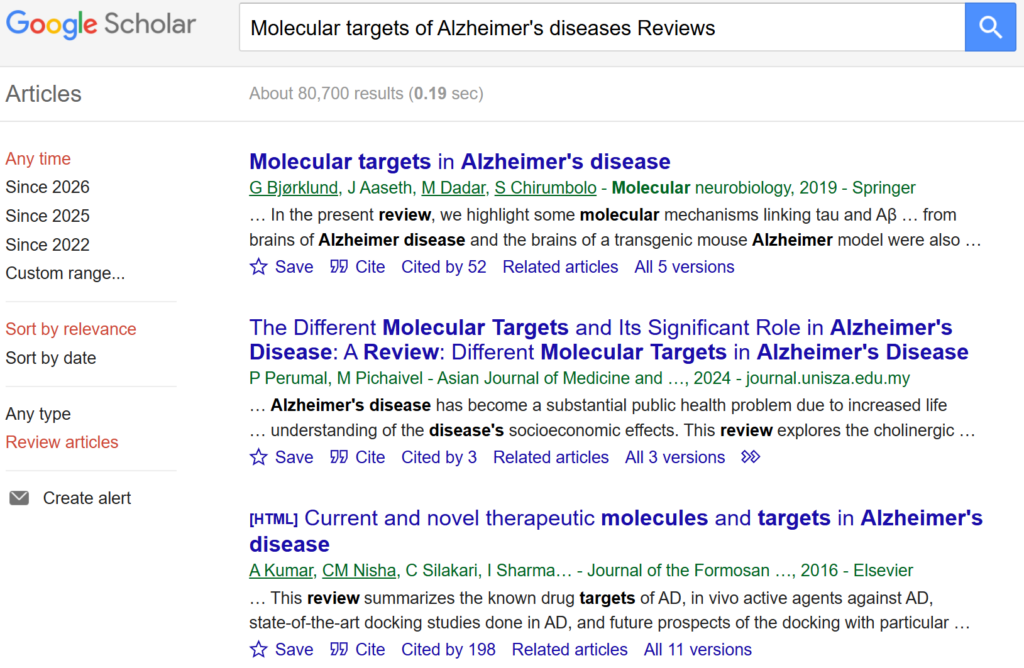
From these articles, compile a list of candidate target proteins repeatedly discussed in the context of disease progression or treatment.
2. Identify Targets Using Omics and Network-Based Approaches
Beyond literature review, computational biology offers data-driven methods to identify potential targets.
Common Approaches
- Differentially Expressed Genes (DEG) analysis
- Protein–protein interaction (PPI) networks
- Pathway enrichment analysis
These methods help identify proteins that:
- Are dysregulated in disease conditions
- Occupy central positions in biological networks
- Influence multiple downstream pathways
Proteins with high network connectivity or regulatory importance are often strong candidates for drug discovery.
3. Reverse Target Identification from Known Compounds
If your research begins with candidate compounds rather than a disease:
- Tools such as SwissTargetPrediction can predict potential protein targets based on chemical similarity and known bioactivity data.
- This approach is useful for:
- Natural products
- Repurposed drugs
- Lead compounds from experimental studies
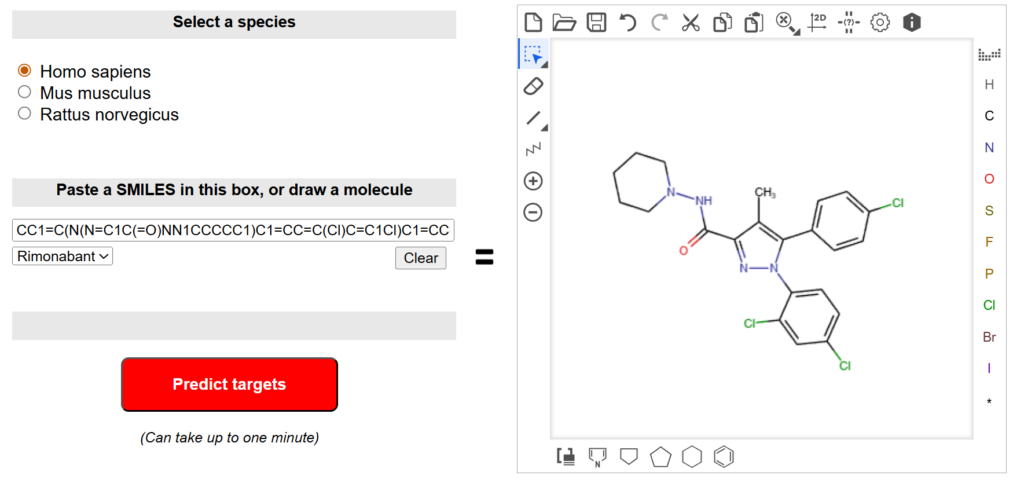
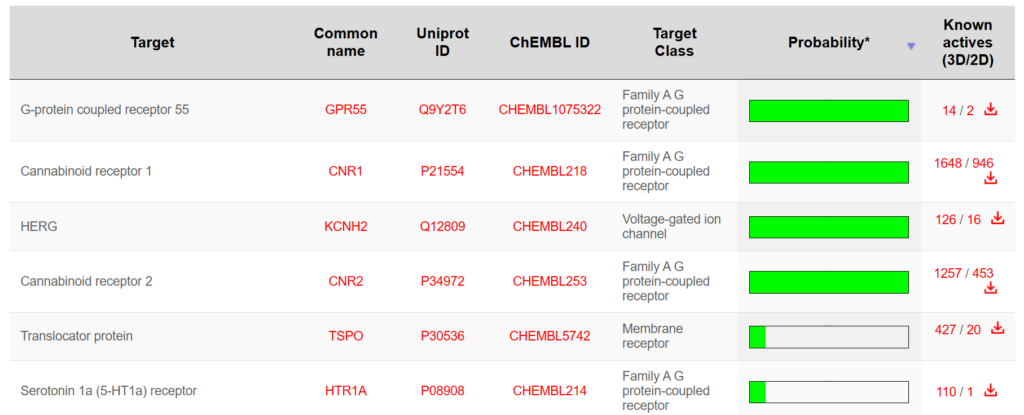
Predicted targets can then be evaluated further using structural and biological criteria.
4. Leverage Existing Laboratory Expertise
If your laboratory already works with a specific protein:
- Selecting that protein as a computational target can be highly effective.
- Advantages include:
- Existing biological knowledge
- Availability of experimental data
- Clear biological relevance
Computational studies can complement wet-lab experiments by offering mechanistic insights and guiding hypothesis generation.
5. Evaluate Structural Availability of the Target Protein
Once candidate proteins are identified, structural feasibility must be assessed.
Key Questions to Ask
- Are high-quality crystal structures available in the Protein Data Bank (PDB)?
- Is the resolution sufficient for docking or simulations?
- Are protein–ligand complex structures available?
Protein–ligand complexes are especially valuable because they:
- Provide binding site information
- Serve as controls for molecular docking
- Enable validation of computational protocols
6. Check for Known Inhibitors or Control Compounds
Before proceeding, confirm whether:
- Known inhibitors or modulators exist
- Experimental binding data is available
These compounds are essential as positive controls in:
- Molecular docking
- Molecular dynamics simulations
- Binding free energy calculations
Controls increase confidence in computational results.
7. Consider Predicted and Modeled Structures
If experimental structures are unavailable:
Predicted Structures
- AlphaFold provides high-quality predicted protein structures.
- Always inspect:
- Confidence scores (pLDDT)
- Disordered regions
- Domain completeness
Homology Modeling
- If related proteins with known structures exist:
- Homology modeling (e.g., via SWISS-MODEL) may be feasible
- Model quality should be carefully assessed before use.
Summary
A suitable target protein for computational drug discovery should satisfy both biological relevance and structural feasibility. A systematic selection process typically includes:
- Disease-focused literature review
- Network or omics-based analysis
- Compound-driven target prediction (if applicable)
- Structural availability and quality checks
- Presence of known inhibitors or control ligands
What’s Next?
Upcoming tutorials from Idgence Research will cover:
- Practical literature mining strategies
- Hands-on DEG and network analysis
- Structural quality assessment for docking
- Preparing target proteins for molecular simulations
Explore related resources:
- Training: https://idgence.org/training
- Research Highlights: https://idgence.org/research
- People: https://idgence.org/people
- Contact: https://idgence.org/contact